Single-nucleus analysis of accessible chromatin in developing mouse forebrain reveals cell-type-specific transcriptional regulation. Integrative single-cell analysis of transcriptional and epigenetic states in the human adult brain. Multiplex single-cell profiling of chromatin accessibility by combinatorial cellular indexing. Simultaneous epitope and transcriptome measurement in single cells. Massively parallel single-nucleus RNA-seq with DroNc-seq. Massively parallel digital transcriptional profiling of single cells. Highly parallel genome-wide expression profiling of individual cells using nanoliter droplets. Droplet barcoding for single-cell transcriptomics applied to embryonic stem cells. Single-cell profiling of the developing mouse brain and spinal cord with split-pool barcoding. Comprehensive single-cell transcriptional profiling of a multicellular organism. Seq-Well: portable, low-cost RNA sequencing of single cells at high throughput. Mapping the mouse cell atlas by Microwell-Seq. Single-cell epigenomics: recording the past and predicting the future. Revealing the vectors of cellular identity with single-cell genomics. Scaling single-cell genomics from phenomenology to mechanism. scGESTALT provides a scalable platform to map lineage relationships between cell types in any system that permits genome editing during development, regeneration, or disease. Generating transgenic lines takes 6 months, and performing barcode editing and generating single-cell libraries involve 7 d of hands-on time. Here, we provide details for (i) generating transgenic zebrafish (ii) performing multi-timepoint barcode editing (iii) building scRNA-seq libraries from brain tissue and (iv) concurrently amplifying lineage barcodes from captured single cells. The recorded lineages are captured, along with thousands of cellular transcriptomes, to build lineage trees with hundreds of branches representing relationships among profiled cell types. The technique generates edits in the barcode array over multiple timepoints using Cas9 and pools of single-guide RNAs (sgRNAs) introduced during early and late zebrafish embryonic development, which distinguishes it from similar Cas9 lineage-tracing methods. We recently established a method, scGESTALT, that combines cumulative editing of a lineage barcode array by CRISPR–Cas9 with large-scale transcriptional profiling using droplet-based single-cell RNA sequencing (scRNA-seq). concentration measurements in UV light) the colorless FastDigest Buffer is recommended.įor methylation sensitivity, refer to product specifications.Lineage relationships among the large number of heterogeneous cell types generated during development are difficult to reconstruct in a high-throughput manner. For applications that require product analysis by fluorescence excitation (e.g. Note: The FastDigest Green Buffer offers the same high performance in DNA digestion and downstream applications as the colorless FastDigest Buffer. Restriction fragment length polymorphism (RFLP).100% buffer compatibility with downstream applications.100% activity of all FastDigest enzymes in the universal buffer.Short incubation times and optimal composition of the universal FastDigest Buffer eliminate star activity effects. 
Therefore, enzymes for downstream applications can be directly added to the FastDigest reaction mix.įor additional convenience FastDigest Green Buffer includes a density reagent and two tracking dyes for direct loading of digestion reaction products on gels. 
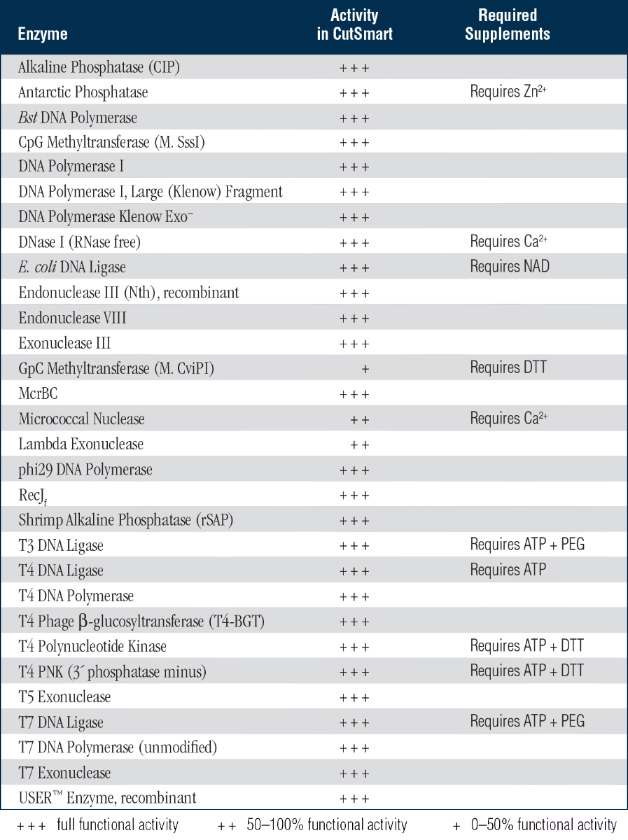
DNA modifying enzymes, such as Klenow Fragment, T4 DNA Ligase, alkaline phosphatases and T4 DNA Polymerase all have 100% activity in FastDigest Buffer. See Reaction Conditions for FastDigest Enzymes for a table of enzyme activity, heat inactivation and incubation times for this and other FastDigest restriction enzymes. The universal buffer allows rapid single-, double-, or multiple DNA digestion within 5–15 minutes eliminating any need for buffer change or subsequent DNA clean-up steps. Thermo Scientific FastDigest NotI is one of an advanced line of fast restriction enzymes that are all 100% active in the universal FastDigest and FastDigest Green reaction buffers. Thermo Scientific FastDigest NotI restriction enzyme recognizes GC^GGCCGC site and cuts best at 37☌ in 5–15 minutes using universal FastDigest Buffer.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
